Recombinant DNA (rDNA) technology /Genetic modification (GM)
Recombinant DNA technology (rDNA) involves the manipulation of an organism’s DNA by cutting it and recombining it in order to change the characteristics of an organism. This can be:
- To introduce a new trait by inserting DNA from another organism or,
- To remove an undesirable characteristic by preventing the expression of a gene.
Introduction of a foreign gene using rDNA technology involves a number of steps as follows:
- An organism that carries the gene of interest is identified: The desired gene can be sourced from any organism including bacteria, fungi, plant or animal species. Unlike conventional breeding methods, rDNA technology presents the opportunity of breaking species boundaries. This means genes can be transferred even between non-related species where this would be near impossible with conventional techniques.
- Isolation and cloning of the gene of interest: Following identification of a donor that carries the gene of interest, DNA is extracted from the individual and the gene of interest isolated. Genes are isolated using enzymes called restriction enzymes. These are enzymes that recognize specific sequences of DNA and cut the molecule at those sequences. Once the gene has been isolated, multiple copies are then produced (i.e. the gene is cloned).
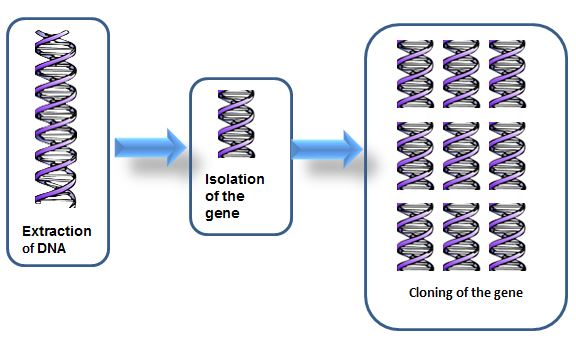
- A transgene construct is developed to include the following:
i. A promoter sequence: This acts as an “on switch” indicating to the machinery of the cell where and when the gene will be expressed. It also acts as a regulator determining how much protein will be produced (expression). The most widely used promoter to date is CaMV35S obtained from the Cauliflower Mosaic Virus. This promoter enables high levels of expression in all plant cells all the time. Another promoter that has been used is Phosphoenolpyruvate (PEP) carboxylase promoter. This promoter only gives expression in the leaves.
ii. The coding region / gene of interest: The coding sequence determines what protein (product) will be produced. This is determined by the sequence of nucleotide bases in the DNA.
iii. A terminator sequence: This acts as an “off switch” indicating to the machinery of the cell that the end of the gene has been reached.
iv. A marker gene: A marker gene makes it easy to identify individuals that have successfully taken up the gene i.e. transformed. Traditionally, antibiotic resistance has been used as a marker. The most widely used markers genes are for antibiotic resistance. Examples here include Kanamycin and Neomycin resistance genes.
Another marker gene that has been used is the luciferase gene. This gene is isolated from insects that glow in the dark. In this case, individuals that have successfully been transformed would show this trait.
- Introduction of the gene into the desired organism using a suitable method:
A number of methods can be used to introduce the transgene construct into the recipient cell. These include:
i. Use of biolistics/ particle bombardment: The transgene construct can be coated with a heavy metal such as tungsten and shot into the recipient cell using a biolistics gun.
ii. Agrobacterium mediated transfer: A naturally occurring bacterium called Agrobacterium tumefaciens can be used. This bacterium is known to infect plants causing them to develop tumors called crown galls. The reason it does this is that it contains a tumour inducing plasmid (Ti plasmid). To use A. tumefaciens for gene transfer, the virulent part of the plasmid is disarmed and replaced with the transgene then allowed to “infect” the plants.
iii. Microinjection: This refers to direct injection of DNA into cells using a fine pipette.
- Selection of transformed individuals: Individuals in which the transgene has been successfully integrated will express the trait associated with the marker gene. These will then be selected for use in a breeding programme to integrate the gene through conventional crossing and backcrossing.
